Structural evolution of an amphibian-specific globin: a computational evolutionary biochemistry approach
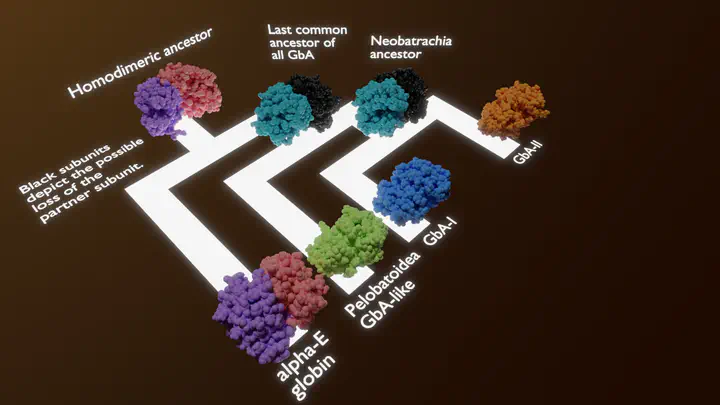 Image credit: Pedro Queiroz
Image credit: Pedro Queiroz
Abstract
Studies on the globin family are continuously revealing insights into the mechanisms of gene and protein evolution. The rise of a new globin gene type in Pelobatoidea and Neobatrachia (Amphibia:Anura) from an $\alpha$-globin precursor provides the opportunity to investigate the genetical and physical mechanisms underlying the origin of new protein structural and functional properties. This amphibian-specific globin (globin A/GbA) discovered in the heart of Rana catesbeiana is a monomer. As the ancestral oligomeric state of $\alpha$-globins is a homodimer, we inferred that the ancestral state was lost somewhere in the GbA lineage. Here, we combined computational molecular evolution with structural bioinformatics to determine how pervasive is the loss of the homodimeric state in the GbA clade. We also characterized the loci of GbA genes in Bufo bufo. We found two GbA clades in Neobatrachia. One of them was deleted in Ranidae but retained and expanded to yield a new globin cluster in Bufonidae species. The loss of the ancestral oligomeric state seems to be pervasive in the GbA clade. However, a taxonomic sampling including more Pelobatoidea as well as early Neobatrachia lineages would be necessary to determine the oligomeric state of the last common ancestor of all GbA. The evidence presented here points out a possible loss of oligomerization in Pelobatoidea GbA due to amino acid substitutions that weaken the homodimeric state. In contrast, the loss of oligomerization in both Neobatrachia GbA clades was linked to independent deletions that disrupted many packing contacts at the homodimer interface.