ggchemplot: an R package for publication-quality 2D chemical structures
This week I faced a situation that many biochemists should relate: preparing figures for a paper in which you have to draw chemical structures. The commercial software ChemDraw, initially developed in 1985 (Evans, 2014), still is the gold standard for drawing publication-quality 2D chemical structures. Because the signature is expensive and the free software available did not met my needs, I started to look for alternatives in R, specially as a ggplot2 (Wickham, 2016) extension. I reasoned that such a fundamental kind of visualization in (bio)chemical and medical sciences would count with a nice R package. There was none.
My next move was to load as a dataframe in R a Structure Data File (SDF) of a 2D chemical compound downloaded from PubChem. The file specify atoms and coordinates. That’s all we need for ggplot2. After the initial plot, I realized that I could make something out of it. After four days writing functions and packaging them, today I used it for creating a figure for my manuscript. Here, I will show how my first ggplot2 extension might help you, R user.
The ggchemplot R package
The workflow of ggchemplot follows three steps:
- Parse a SDF file with
ggchemplot1().
- Visualize the preliminary plot and modify the data with helper functions for polishing.
- Plot the final visualization with
ggchemplot2().
Short tutorial
Install the package from my GitHub repository.
# Install the remotes package if not already installed
install.packages("remotes")
# Install ggchemplot from GitHub
remotes::install_github("JPFQueiroz/ggchemplot")The input file in this example (P1_pyridone.sdf) was prepared through the drawing tool of PubChem. Other R users might also prepare the input structure directly from R with rcdk (Guha, 2007). Use ggchemplot1 to parse the file and generate the initial data object.
# Load ggchemplot
library(ggchemplot)
# Parse the data and generate initial plot
p1 <- ggchemplot1(sdf_file = "P1_pyridone.sdf")
# Check the initial plot
p1$plot
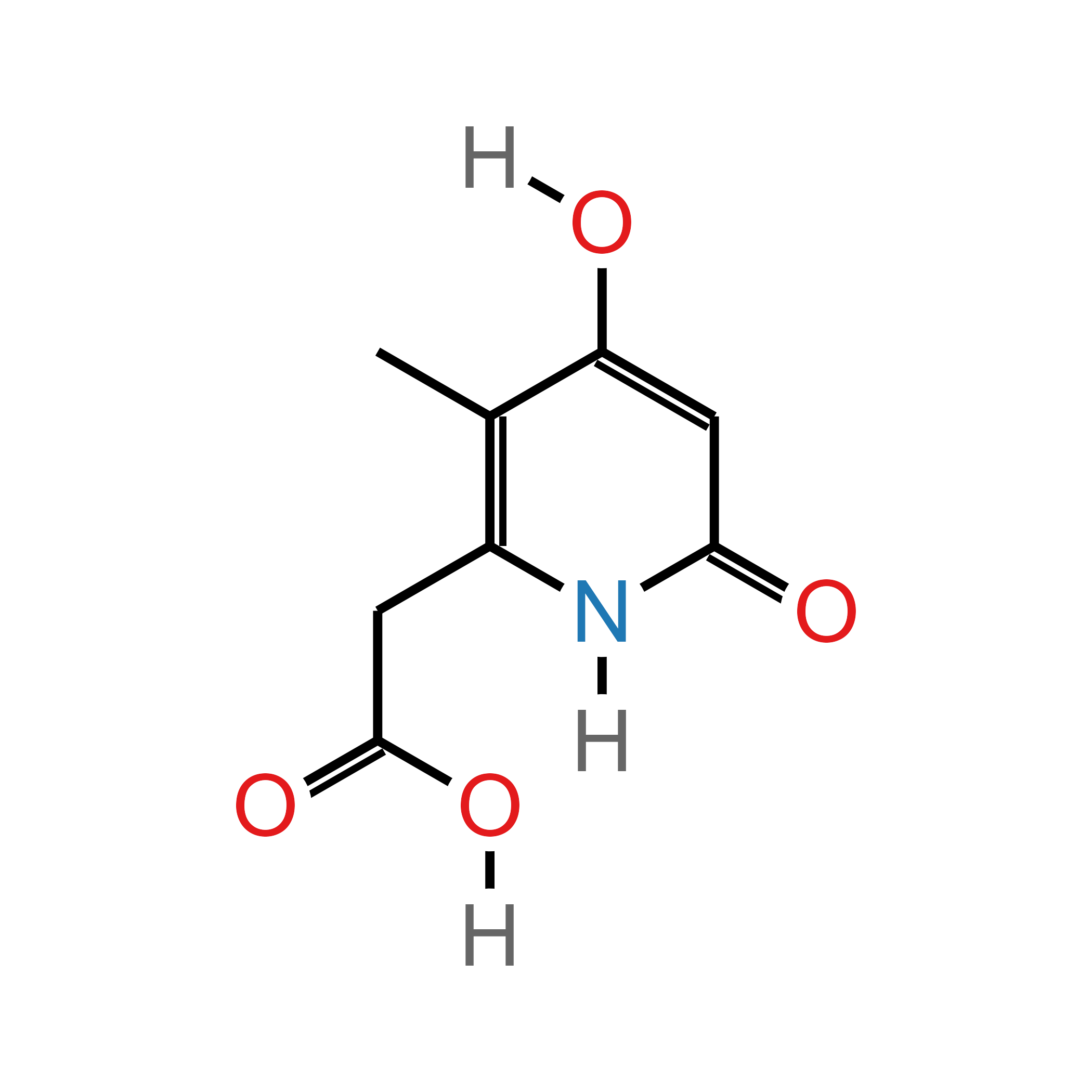
Quickly visualize the atom and bond ids in p1$atoms on the plot with the argument show_ids = TRUE in ggchemplot2(). The ids visualization can guide through the polishing step, where many functions use as argument ids of atoms or bonds.
# Load dplyr to use the pipe operator %>%
library(dplyr)
p1 %>%
ggchemplot2(show_ids = TRUE)
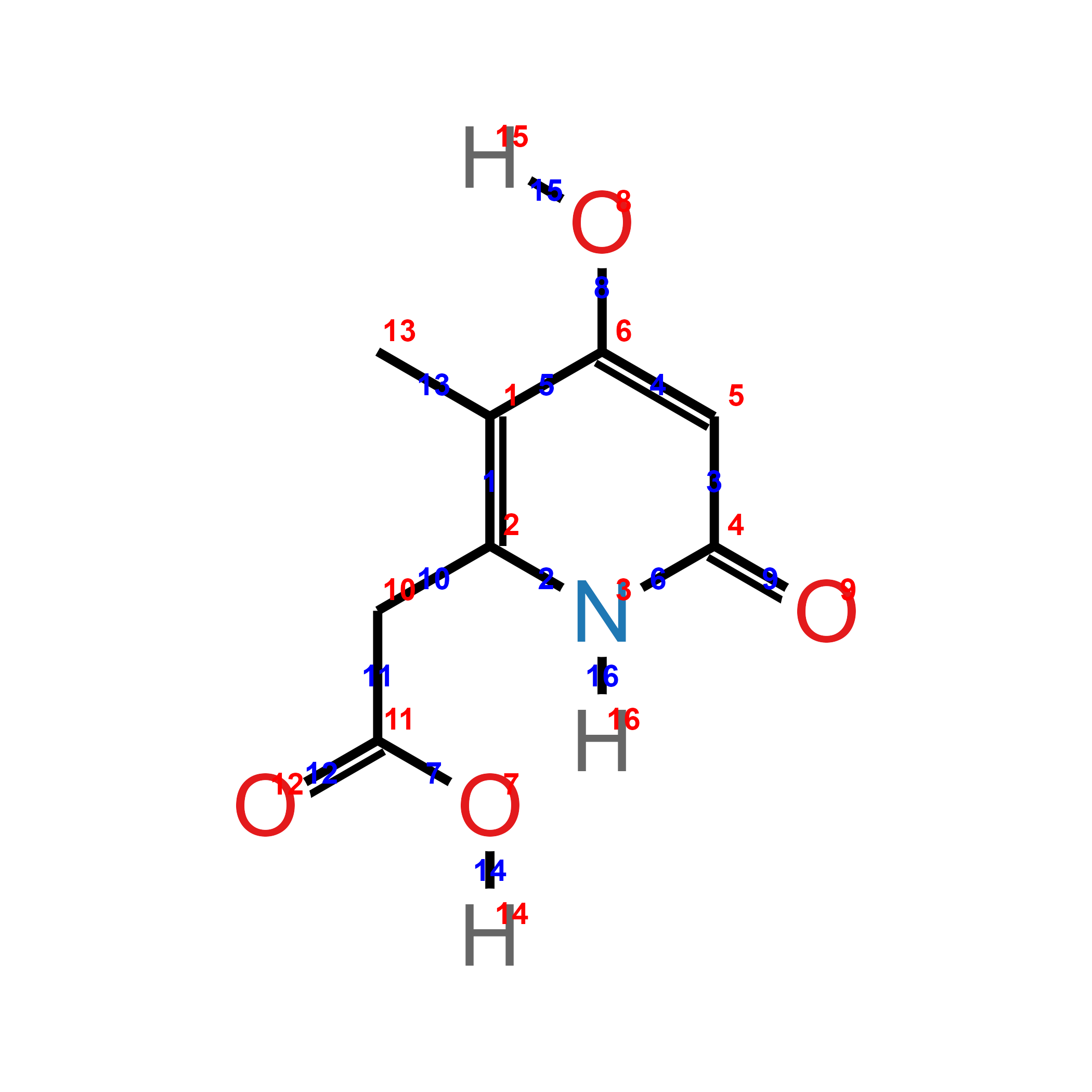
Polish the visualization using the arguments available in ggchemplot1(), modifying the underlying data with helper functions or through manual wrangling of the dataframe objects, and finally to ggchemplot2().
# Publication-quality plot
p1 <- ggchemplot1(
sdf_file = "P1_pyridone.sdf",
title = NULL,
collapse_hydrogens = TRUE,
rotation = 90,
label_padding = 1,
show_atom_circles = TRUE,
hide_carbon_circles = TRUE,
circle_stroke = 0,
show_atom_labels = TRUE,
hide_carbon_labels = TRUE,
bond_width = 2,
atom_size = 18,
label_size = 12,
double_bond_offset = 0.3,
custom_atom_colors = NULL,
paint_it_black = TRUE,
H_offset = c(0.50, 1)) %>%
change_label(atom_id = 7, new_label = "Fe") %>%
add_hashed_bond(from_id = 7, to_id = 11,
wedge_thickness = 0.3,n_hashes = 6,
shorten = c(0.3, 0)) %>%
ggchemplot2()
# Plot
p1
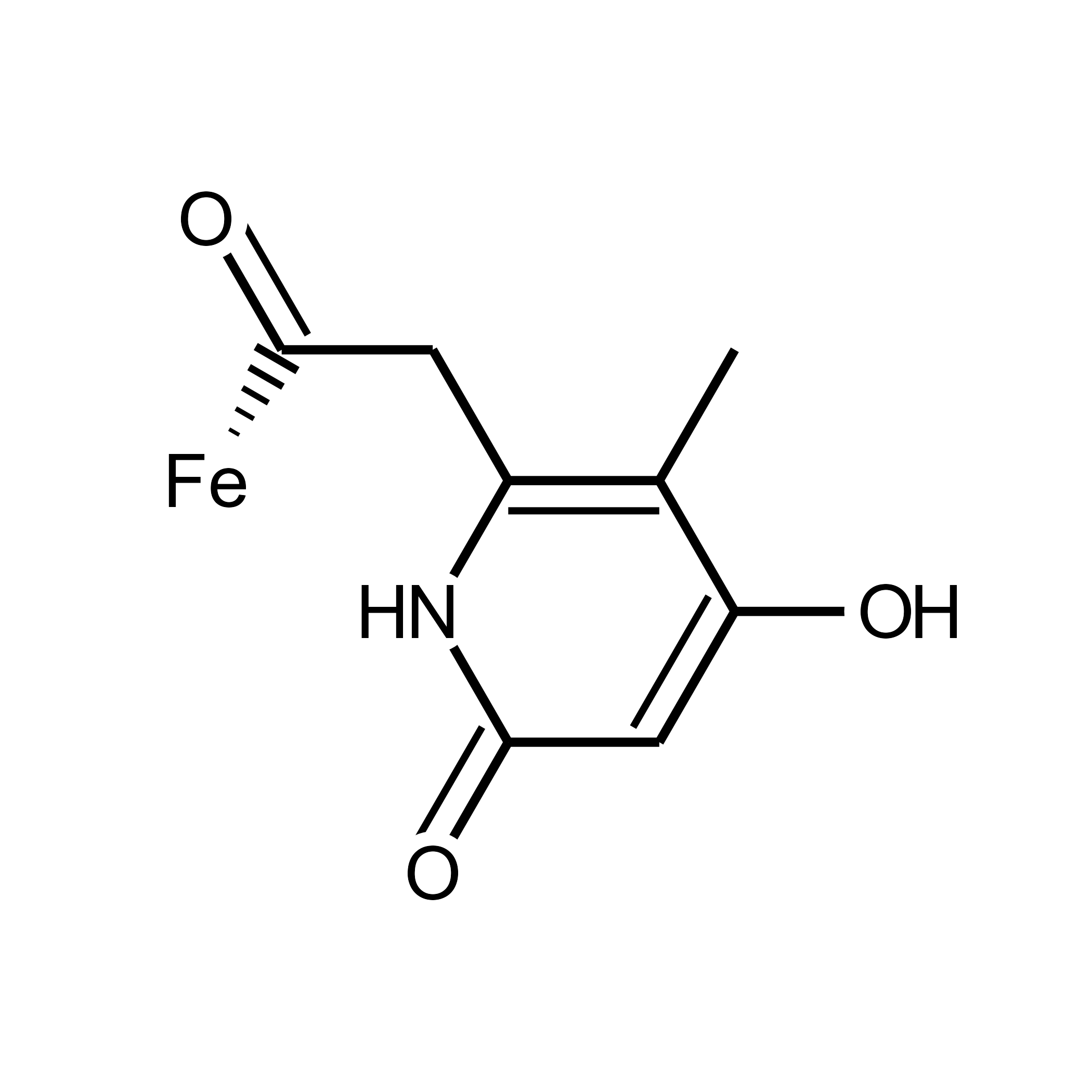
The basic tutorial ends here. You can cite ggchemplot (Fernandes-Queiroz, 2026) in your publications using the BibTex citation below. Your feedback is welcome.
@software{ggchemplot,
author = {João Pedro Fernandes Queiroz},
title = {JPFQueiroz/ggchemplot: ggchemplot: an R package for visualization of chemical structures},
month = may,
year = 2026,
publisher = {Zenodo},
version = {v0.2-beta.1},
doi = {10.5281/zenodo.20088880},
url = {https://doi.org/10.5281/zenodo.20088880},
}